What is the flow of information through the cell? PowerPoint PPT Presentation
1 / 43
Title: What is the flow of information through the cell?
1
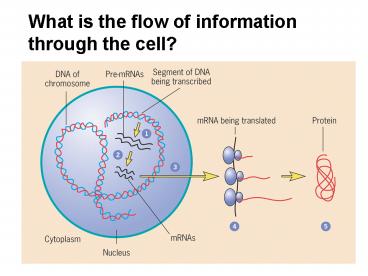
What is the flow of information through the cell?
2
Double helix - antiparallel polymers
Major groove Minor groove
3
5 3
4
Purine Pyrimidine
5
(No Transcript)
6
06_12_asymmetrical.jpg
7
(No Transcript)
8
- Transcription
- dsDNA template
- Nucleotides (ACGU) make ssRNA
- Need to separate strands.
- Nucleotides added to free 3 OH (5?3)
9
Classes of RNA
mRNA
rRNA
tRNA
also snRNA and microRNA
10
Prokaryotes
untranslated regions 5 UTR
3 UTR
coding sequence
promoter (not transcribed)
DNA
mRNA
RNA Polymerase Ribosome
polypeptide
11
Eukaryotes
untranslated regions 5 URT
3 UTR
promoter (not transcribed)
coding sequence
DNA
pre mRNA
mRNA
RNA Polymerase Ribosome
polypeptide
12
Bacterial Promoter Elements
- Transcription start 1
- Consensus sequence 35 TTGACA, recognized by
? - Pribnow box -10, TATAAT determines 1
- Terminator sequence where polymerase stops
13
(No Transcript)
14
Initiation of transcription in prokaryotes
15
Initiation of eukaryotic transcription by RNA
Pol II (mRNA) TF transcription factor (compare
with prokaryotic sigma factor)
16
- Eukaryotic mRNA
- 5 cap,
- 5 UTR
- coding region
- 3 UTR
- 3 poly-A tail
17
mRNA processing in Eukaryotes 5 cap
added Remove 3 end Poly-A tail
added Introns removed Exons joined
18
Gene cloning - making lots of copies...
1. Make a library of small pieces of DNA (2
types) 2. Find the one piece you want 3. Insert
it into a vector 4. Grow it in a new organism
(bacteria, euk. cells)
Isolate DNA, fragment with RE
Isolate mRNA, convert to cDNA with reverse
transcriptase
Genomic library
cDNA library
19
(No Transcript)
20
07_37_Protein.produc.jpg
Overview of gene expression in eukaryotes
21
07_28_ribosome.jpg
22
07_26_2_adaptors.jpg
Two adapters link an amino acid to a codon
23
(No Transcript)
24
07_32_initiation.jpg
Intiation of translation in Eukaryotes
25
07_33_mRNA.encode.jpg
Intiation of translation in Prokaryotes
26
07_30_3_step_cycle.jpg
Elongation of proteins
4
5
27
07_34_stop codon.jpg
Termination of translation
28
Mutations
- Frameshift Adding or removing 1 or 2 nucleotides
results in changes the reading frame from that
point on. - Nonsense Changing an amino acid codon to a stop
codon results in truncated proteins - Missense Changing an amino acid codon to one
encoding a different amino acid - effect depends
on type of amino acid and where in the protein.
29
(No Transcript)
30
04_03_20 amino acids.jpg
31
Side chains interact via all of the noncovalent
bonds
32
Primary structure (1) of a protein Arabidopsis
?-glucosidase (single letter codes) MSSLHWFPNIFIV
VVVFFSLRSSQVVLEEEESTVVGYGYVVRSVGVDSNRQVLTAKLDLIKPS
SVYAPDIKSLNLHVSLETSERLRIRITDSSQQRWEIPETVIPRAGNHSPR
RFSTEEDGGNSPENNFLADPSSDLVFTLHNTTPFGFSVSRRSSGDILFDT
SPDSSDSNTYFIFKDQFLQLSSALPENRSNLYGIGEHTKRSFRLIPGETM
TLWNADTGSENPDVNLYGSHPFYMDVRGSKGNEEAGTTHGVLLLNSNGMD
VKYEGHRITYNVIGGVIDLYVFAGPSPEMVMNQYTELIGRPAPMPYWSFG
FHQCRYGYKNVSDLEYVVDGYAKAGIPLEVMWTDIDYMDGYKDFTLDPVN
FPEDKMQSFVDTLHKNGQKYVLILDPGIGVDSSYGTYNRGMEADVFIKRN
GEPYLGEVWPGKVYFPDFLNPAAATFWSNEIKMFQEILPLDGLWIDMNEL
SNFITSPLSSGSSLDDPPYKINNSGDKRPINNKTVPATSIHFGNISEYDA
HNLYGLLEAKATHQAVVDITGKRPFILSRSTFVSSGKYTAHWTGDNAAKW
EDLAYSIPGILNFGLFGIPMVGADICGFSHDTTEELCRRWIQLGAFYPFA
RDHSSLGTARQELYLWDSVASSARKVLGLRMRLLPHLYTLMYEAHVSGNP
IARPLFFSFPQDTKTYEIDSQFLIGKSIMVSPALKQGAVAVDAYFPAGNW
FDLFNYSFAVGGDSGKHVRLDTPADHVNVHVREGSIVAMQGEALTTRDAR
KTPYQLLVVASRLENISGELFLDDGENLRMGAGGGNRDWTLVKFRCYVTG
KSVVLRSEVVNPEYASKMKWSIGKVTFVGFENVENVKTYEVRTSERLRSP
RISLIKTVSDNDDPRFLSVEVSKLSLLVGKKFEMRLRLT
33
Secondary structure (2) ?-helix H-bonds
between CO and N-H of backbone. (No R-groups
involved)
34
Secondary structure ?-sheet H-bonds between
CO and N-H of backbone. (No R-groups involved)
35
Tertiary structure - the entire polypeptide
?-helix
?-sheet
loops and turns disulfide bridge
ribonuclease
36
Quaternary structure - multiple subunits
e.g. hemoglobin
37
04_20_protein domains.jpg
lactic (lactate) dehydrogenase
immunoglobulin light chain
cytochrome b562
38
Domains - discrete modules within tertiary
structure that fold independently and have a
specific function. 4 domains of phospholipase C
39
Motif - a recurring substructure ?/? barrel
e.g. ?-amylase
40
Motif - a recurring substructure coiled coil
e.g. myosin
41
How do proteins get to their folded state?
unfolded
native conformation
42
04_21_Serine proteases.jpg
Proteins with different functions may have
similar shape - members of a family with a common
ancestor.
43
04_22_protein subunit.jpg