Medical Informatics PowerPoint PPT Presentation
1 / 49
Title: Medical Informatics
1
Medical Informatics
Bioinformatics
2
Mining Bio-Medical Mountains How Computer Science
can help Biomedical Research and Health Sciences
Anil Jegga Division of Biomedical Informatics,
Cincinnati Childrens Hospital Medical Center
(CCHMC) Department of Pediatrics, University of
Cincinnati http//anil.cchmc.org Anil.Jegga_at_cchmc.
org
3
Acknowledgement
- Biomedical Engineering/Bioinformatics
- Jing Chen
- Sivakumar Gowrisankar
- Vivek Kaimal
- Computer Science
- Amit Sinha
- Mrunal Deshmukh
- Divya Sardana
- Electrical Engineering
- Nishanth Vepachedu
4
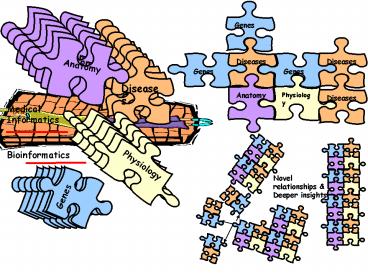
Two Separate Worlds..
Medical Informatics
Bioinformatics the omes
PubMed
Proteome
Disease Database
Patient Records
OMIM Clinical Synopsis
Clinical Trials
382 omes so far and there is UNKNOME too -
genes with no function known http//omics.org/inde
x.php/Alphabetically_ordered_list_of_omics
With Some Data Exchange
5
now. The number 1 FAQ
How much biology should I know??
No simple or straight-forward answer
unfortunately!
But the mantra is Interact routinely with
biologists OR Work with the biologists or the
biological data
6
But I want to learn some basics
- http//www.ncbi.nlm.nih.gov/Education
- http//www.ebi.ac.uk/2can/
- http//www.genome.gov/Education/
- http//genomics.energy.gov/
- Books
- Introduction to Bioinformatics by Teresa Attwood,
David Parry-Smith - A Primer of Genome Science by Gibson G and Muse
SV - Bioinformatics A Practical Guide to the Analysis
of Genes and Proteins, Second Edition by Andreas
D. Baxevanis, B. F. Francis Ouellette - Algorithms on Strings, Trees, and Sequences
Computer Science and Computational Biology by Dan
Gusfield - Bioinformatics Sequence and Genome Analysis by
David W. Mount - Discovering Genomics, Proteomics, and
Bioinformatics by A. Malcolm Campbell and Laurie
J. Heyer
7
And the other FAQs.
- What bioinformatics topics are closest to
computer science? - Should computer science departments involve
themselves in preparing their graduates for
careers in bioinformatics? - And if so, what topics should they cover?
- And how much biology should they be taught?
- Lastly, how much effort should be expended in
re-directing computer scientists to do work in
bioinformatics?
Cohen, 2005 Communications of the ACM
8
Issues to be considered..
- Computer science Vs molecular biology Subject
Scientists - Cultural differences - Current goals of molecular biology, genomics (or
biomedical research in a broader sense) - Data types used in bioinformatics or genomics
- Areas within computer science of interest to
biologists - Bioinformatics research - Employment
opportunities
9
Biological Challenges - Computer Engineers
- Post-genomic Era and the goal of bio-medicine
- to develop a quantitative understanding of how
living things are built from the genome that
encodes them. - Deciphering the genome code
- Identifying unknown genes and assigning function
by computational analysis of genomic sequence - Identifying the regulatory mechanisms
- Identifying their role in normal
development/states vs disease states
10
Biological Challenges - Computer Engineers
- Data Deluge exponential growth of data silos and
different data types - Human-computer interaction specialists need to
work closely with academic and clinical
biomedical researchers to not only manage the
data deluge but to convert information into
knowledge. - Biological data is very complex and interlinked!
- Creating information systems that allow
biologists to seamlessly follow these links
without getting lost in a sea of information - a
huge opportunity for computer scientists.
11
Biological Challenges - Computer Engineers
A major goal in molecular biology is Functional
Genomics Study of the relationships among genes
in DNA their function in normal and disease
states
- Networks, networks, and networks!
- Each gene in the genome is not an independent
entity. Multiple genes interact to perform a
specific function. - Environmental influences Genotype-environment
interaction - Integrating genomic and biochemical data together
into quantitative and predictive models of
biochemistry and physiology - Computer scientists, mathematicians, and
statisticians will ALL be an integral and
critical part of this effort.
12
Informatics Biologists Expectations
- Representation, Organization, Manipulation,
Distribution, Maintenance, and Use of
information, particularly in digital form. - Functional aspect of bioinformatics
Representation, Storage, and Distribution of
data. - Intelligent design of data formats and databases
- Creation of tools to query those databases
- Development of user interfaces or visualizations
that bring together different tools to allow the
user to ask complex questions or put forth
testable hypotheses.
13
Informatics Biologists Expectations
- Developing analytical tools to discover knowledge
in data - Levels at biological information is used
- comparing sequences predict function of a newly
discovered gene - breaking down known 3D protein structures into
bits to find patterns that can help predict how
the protein folds - modeling how proteins and metabolites in a cell
work together to make the cell function.
14
Finally.What does informatics mean to
biologists?
- The ultimate goal of analytical bioinformaticians
is to develop predictive methods that allow
biomedical researchers and scientists to model
the function and phenotype of an organism based
only on its genomic sequence. This is a grand
goal, and one that will be approached only in
small steps, by many scientists from different
but allied disciplines working cohesively.
15
Biology Data Structures
- Four broad categories
- Strings To represent DNA, RNA, amino acid
sequences of proteins - Trees To represent the evolution of various
organisms (Taxonomy) or structured knowledge
(Ontologies) - Sets of 3D points and their linkages To
represent protein structures - Graphs To represent metabolic, regulatory, and
signaling networks or pathways
16
Biology Data Structures
- Biologists are also interested in
- Substrings
- Subtrees
- Subsets of points and linkages, and
- Subgraphs.
Beware Biological data is often characterized by
huge size, the presence of laboratory errors
(noise), duplication, and sometimes unreliability.
17
Support Complex Queries A typical demand
- Get me all genes involved in or associated with
brain development that are differentially
expressed in the Central Nervous System. - Get me all genes involved in brain development in
human and mouse that also show iron ion binding
activity. - For this set of genes, what aspects of function
and/or cellular localization do they share? - For this set of genes, what mutations are
reported to cause pathological conditions?
18
Model Organism Databases Common Issues
- Heterogeneous Data Sets - Data Integration
- From Genotype to Phenotype
- Experimental and Consensus Views
- Incorporation of Large Datasets
- Whole genome annotation pipelines
- Large scale mutagenesis/variation projects
(dbSNP) - Computational vs. Literature-based Data
Collection and Evaluation (MedLine) - Data Mining
- extraction of new knowledge
- testable hypotheses (Hypothesis Generation)
19
Bioinformatic Data-1978 to present
- DNA sequence
- Gene expression
- Protein expression
- Protein Structure
- Genome mapping
- SNPs Mutations
- Metabolic networks
- Regulatory networks
- Trait mapping
- Gene function analysis
- Scientific literature
- and others..
20
Human Genome Project Data Deluge
No. of Human Gene Records currently in NCBI
29413 (excluding pseudogenes, mitochondrial genes
and obsolete records). Includes 460 microRNAs
NCBI Human Genome Statistics as on February12,
2008
21
The Gene Expression Data Deluge
Till 2000 413 papers on microarray!
Problems Deluge! Allison DB, Cui X, Page GP,
Sabripour M. 2006. Microarray data analysis from
disarray to consolidation and consensus. Nat Rev
Genet. 7(1) 55-65.
22
Information Deluge..
- 3 scientific journals in 1750
- Now - gt120,000 scientific journals!
- gt500,000 medical articles/year
- gt4,000,000 scientific articles/year
- gt16 million abstracts in PubMed derived from
gt32,500 journals
23
Data-driven Problems..
- Generally, the names refer to some feature of the
mutant phenotype - Dickies small eye (Thieler et al., 1978, Anat
Embryol (Berl), 155 81-86) is now Pax6 - Gleeful "This gene encodes a C2H2 zinc finger
transcription factor with high sequence
similarity to vertebrate Gli proteins, so we have
named the gene gleeful (Gfl)." (Furlong et al.,
2001, Science 293 1632)
Whats in a name!
Rose is a rose is a rose is a rose!
Gene Nomenclature
- Disease names
- Mobius Syndrome with Polands Anomaly
- Werners syndrome
- Downs syndrome
- Angelmans syndrome
- Creutzfeld-Jacob disease
- Accelerin
- Antiquitin
- Bang Senseless
- Bride of Sevenless
- Christmas Factor
- Cockeye
- Crack
- Draculin
- Dickies small eye
- Draculin
- Fidgetin
- Gleeful
- Knobhead
- Lunatic Fringe
- Mortalin
- Orphanin
- Profilactin
- Sonic Hedgehog
24
Rose is a rose is a rose is a rose.. Not Really!
What is a cell?
- any small compartment
- (biology) the basic structural and functional
unit of all organisms they may exist as
independent units of life (as in monads) or may
form colonies or tissues as in higher plants and
animals - a device that delivers an electric current as a
result of chemical reaction - a small unit serving as part of or as the nucleus
of a larger political movement - cellular telephone a hand-held mobile
radiotelephone for use in an area divided into
small sections, each with its own short-range
transmitter/receiver - small room in which a monk or nun lives
- a room where a prisoner is kept
Image Sources Somewhere from the internet
25
Foundation Model Explorer
26
- COLORECTAL CANCER 3-BP DEL, SER45DEL
- COLORECTAL CANCER SER33TYR
- PILOMATRICOMA, SOMATIC SER33TYR
- HEPATOBLASTOMA, SOMATIC THR41ALA
- DESMOID TUMOR, SOMATIC THR41ALA
- PILOMATRICOMA, SOMATIC ASP32GLY
- OVARIAN CARCINOMA, ENDOMETRIOID TYPE, SOMATIC
SER37CYS - HEPATOCELLULAR CARCINOMA SOMATIC SER45PHE
- HEPATOCELLULAR CARCINOMA SOMATIC SER45PRO
- MEDULLOBLASTOMA, SOMATIC SER33PHE
The REAL Problems
Many disease states are complex, because of many
genes (alleles ethnicity, gene families, etc.),
environmental effects (life style, exposure,
etc.) and the interactions.
27
The REAL Problems
28
Methods for Integration
- Link driven federations
- Explicit links between databanks.
- Warehousing
- Data is downloaded, filtered, integrated and
stored in a warehouse. Answers to queries are
taken from the warehouse. - Others.. Semantic Web, etc
29
Link-driven Federations
- Creates explicit links between databanks
- query get interesting results and use web links
to reach related data in other databanks - Examples NCBI-Entrez, SRS
30
http//www.ncbi.nlm.nih.gov/Database/datamodel/
31
http//www.ncbi.nlm.nih.gov/Database/datamodel/
32
http//www.ncbi.nlm.nih.gov/Database/datamodel/
33
http//www.ncbi.nlm.nih.gov/Database/datamodel/
34
http//www.ncbi.nlm.nih.gov/Database/datamodel/
35
Link-driven Federations
- Advantages
- complex queries
- Fast
- Disadvantages
- require good knowledge
- syntax based
- terminology problem not solved
36
Data Warehousing
Data is downloaded, filtered, integrated and
stored in a warehouse. Answers to queries are
taken from the warehouse.
- Advantages
- Good for very-specific, task-based queries and
studies. - Since it is custom-built and usually
expert-curated, relatively less error-prone.
- Disadvantages
- Can become quickly outdated needs constant
updates. - Limited functionality For e.g., one
disease-based or one system-based.
37
Algorithms in Bioinformatics
- Finding similarities among strings
- Detecting certain patterns within strings
- Finding similarities among parts of spatial
structures (e.g. motifs) - Constructing trees
- Phylogenetic or taxonomic trees evolution of an
organism - Ontologies structured/hierarchical
representation of knowledge - Classifying new data according to previously
clustered sets of annotated data
38
Algorithms in Bioinformatics
- Reasoning about microarray data and the
corresponding behavior of pathways - Predictions of deleterious effects of changes in
DNA sequences - Computational linguistics NLP/Text-mining.
Published literature or patient records - Graph Theory Gene regulatory networks,
functional networks, etc. - Visualization and GUIs (networks, application
front ends, etc.)
39
Disease Gene Identification and Prioritization
Hypothesis Majority of genes that impact or
cause disease share membership in any of several
functional relationships OR Functionally similar
or related genes cause similar phenotype.
- Functional Similarity Common/shared
- Gene Ontology term
- Pathway
- Phenotype
- Chromosomal location
- Expression
- Cis regulatory elements (Transcription factor
binding sites) - miRNA regulators
- Interactions
- Other features..
40
Background, Problems Issues
- Most of the common diseases are multi-factorial
and modified by genetically and mechanistically
complex polygenic interactions and environmental
factors. - High-throughput genome-wide studies like linkage
analysis and gene expression profiling, tend to
be most useful for classification and
characterization but do not provide sufficient
information to identify or prioritize specific
disease causal genes.
41
Background, Problems Issues
- Since multiple genes are associated with same or
similar disease phenotypes, it is reasonable to
expect the underlying genes to be functionally
related. - Such functional relatedness (common pathway,
interaction, biological process, etc.) can be
exploited to aid in the finding of novel disease
genes. For e.g., genetically heterogeneous
hereditary diseases such as Hermansky-Pudlak
syndrome and Fanconi anaemia have been shown to
be caused by mutations in different interacting
proteins.
42
PPI - Predicting Disease Genes
- Direct proteinprotein interactions (PPI) are one
of the strongest manifestations of a functional
relation between genes. - Hypothesis Interacting proteins lead to same or
similar disease phenotypes when mutated. - Several genetically heterogeneous hereditary
diseases are shown to be caused by mutations in
different interacting proteins. For e.g.
Hermansky-Pudlak syndrome and Fanconi anaemia.
Hence, proteinprotein interactions might in
principle be used to identify potentially
interesting disease gene candidates.
43
- Prioritize candidate genes in the interacting
partners of the disease-related genes - Training sets disease related genes
- Test sets interacting partners of the training
genes
44
- Example Breast cancer
15
342
2469
45
ToppGene General Schema
46
(No Transcript)
47
(No Transcript)
48
PubMed
OMIM
49
http//sbw.kgi.edu/