35 Proofreading Repair by Klenow Fragment of DNA pol I - PowerPoint PPT Presentation
1 / 19
Title:
35 Proofreading Repair by Klenow Fragment of DNA pol I
Description:
However, pol III in a multi-protein aggregate termed the ... on lagging strand must bind and dissociate constantly, yet keep at same pace as leading strand ... – PowerPoint PPT presentation
Number of Views:1095
Avg rating:3.0/5.0
Title: 35 Proofreading Repair by Klenow Fragment of DNA pol I
1
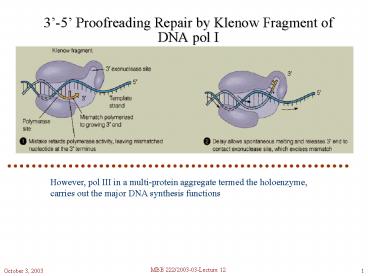
3-5 Proofreading Repair by Klenow Fragment of
DNA pol I
However, pol III in a multi-protein aggregate
termed the holoenzyme, carries out the major DNA
synthesis functions
2
DNA Polymerase III Holoenzyme
? - PolC gene product.- Part of core polymerase.
It provides the actual polymerase activity.. ? -
Part of the core polymerase. Contains a 3'-5'
exonuclease activity (proof reading). ? - Part
of the core polymerase .Unknown function. ? -
This dimeric protein dimerizes the holoenzyme,
holding leading and lagging strand polymerases
together so both DNA strands are elongated at the
replication fork ? - Mediates the switch from
making RNA primers (with primase) to making DNA
(with PolC) ? - Functions as a sliding clamp to
hold the holoenzyme complex to DNA, making the
enzyme very processive (stays on the DNA for the
addition of thousands of nucleotides) ?, ?',
?, ? ,? - This group of proteins is called the ?
complex (also called the clamp loader). It is
composed of one copy of all the proteins except
?, of which there are 2 or 3 copies. The clamp
loader wraps the ? clamp onto the DNA.
3
DNA Polymerase III Holoenzyme
Unknown fxn
3?5 exonuclease
Pol C polymerase
Sliding Clamp
Dimerizing Units
the clamp loader
4
Structure of the ? clamp
Crystal Structure Of The Processivity Clamp Gp45
From Bacteriophage T4
5
How does the ? clamp get wrapped around the DNA?
-a conformational change driven by ATP binding
leads complex to bind DNA and g to bring about
ring opening - hydrolysis of ATP required to
alter the stable structure of the ? clamp and
close ring -happens only once per round of
replication on leading strand, while on lagging
strand must bind and dissociate constantly, yet
keep at same pace as leading strand
6
Eukaryotic DNA Polymerases
Distinguished from each other based on
intracellular locations, kinetic properties, and
responses to inhibitors
(I)
(III)
(II)
7
DNA Replication Fork
- many other proteins involved in DNA replication
than just the polymerases. - these other proteins
perform a variety of tasks related to the
replication of DNA
Figure 24.6
8
Other Proteins Involved in Replication
DNA Ligase Covalently closes nicks in double
stranded DNA nick must contain 3hydroxyl and
5phosphoryl termini Primase - active only in
the presence of other proteins (in complex called
theprimosome), including helicase -
synthesizes RNA primers
9
- Helicases
- - enzymes that use ATP to actively unwind DNA
- Polymerase Accessory Proteins
- - help to keep DNA pol III processive (i.e.
moving continuously on the same strand of DNA,
instead of coming on and off). - Single Stranded DNA Binding Proteins (SSB)
- promote the denaturation of DNA by binding
co-operatively to ss template and maintaining it
in an extended ss conformation - Topoisomerases
- - proteins that relieve super-helical stress
10
Figure 24.26 Action of SSBs
Figure 24.27 a model for helicase action
11
Topoisomerases
Enzymes that catalyze interconversion of
topoisomers of DNA (Change the Super-Helicity)
Two Types Topoisomerase I - relaxes negatively
supercoiled DNA - passively Done. Driven by
release of energy of supercoiling - nicks
(breaks) one strand the other strand swivels
around it. (Broken strand then sealed)
Topoisomerase II - can relax supercoiled DNA or
introduce supercoils - hydrolyzes ATP - breaks
both strands and seals them
12
Topoisomerase I
13
Topoisomerase II
The action of topoisomerase II on DNA results in
an increase or a decrease in the linking number
by ( or -) 2
14
Interconversions catalyzed by Topoisomerase II
Knots
15
Topoisomerase II
16
Action of the E.coli DNA uracil repair system (a
specific type of Base Excision Repair)
Uracil-DNA N-glycosylase is the key enzyme
Uracil is not normally found in DNA and can be
removed by a specific repair process Question
since U properly base pairs with A, why bother to
remove it from DNA (i.e. there would be no
mutation)? Answer the target of this repair
system is probably U derived from deamination of
C, rather than U that was misincorporated during
DNA synthesis the former would cause a mutation
(GC to AT) (more later)
17
DNA Information Restructuring
Different from DNA replication Replication A
major metabolic event highly efficient
effective enzymes other proteins
involved Restructuring Restructuring is a broad
term which includes several different
processes Quantitatively minor pathways, but very
important
18
Types of Restructuring Processes
- Restriction Modification
- Protective mechanism for prokaryotic cells.
(Restriction Enzymes)-associated with
modification (i.e. methylation via methylase)
function-can be associated with the same or with
a separate enzyme - Metabolic Responses to DNA damage
- Mutagenesis
- Repair
- Recombination
- Contents of genome shuffled- occurs mostly
during sexual reproduction - Gene/DNA Rearrangements
- Transposition and chromosomal Integration
- Gene Amplification
19
1. Restriction Modification
Figure 25.5 Host-induced restriction and
modification.
Sometimes Bacteriophage Lambda can escape
restriction by E. coli (host) endonucleases this
occurs when its DNA is modified by the host
methylase